Kinases are a class of enzymes that are responsible for transferring the main chemical energy source used by the body’s cells. As such, they play important roles in diverse cellular processes, including signaling, differentiation, proliferation and metabolism. But since they are so ubiquitous, mutated versions of kinases are frequently found in cancers. Many cancer treatments involve targeting these mutant kinases with specific inhibitors.
Understanding the exact genetic mutations that lead to these aberrant kinases can therefore be critical in predicting the progression of a given patient’s cancer and tailoring the appropriate response.
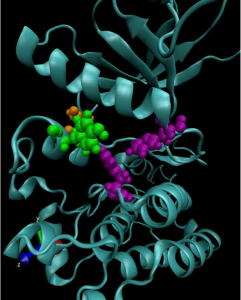
To achieve this understanding on a more fundamental level, a team of researchers from the University of Pennsylvania’s School of Engineering and Applied Science and Perelman School of Medicine, the Children’s Hospital of Philadelphia (CHOP) and researchers at the Yale School of Medicine’s Cancer Biology Institute, have constructed molecular simulations of a mutant kinase implicated in pediatric neuroblastoma, a childhood cancer impacting the central nervous system.
Using their computational model to study the relationship between single-point changes in the kinase’s underlying gene and the altered structure of the protein it ultimately produces, the researchers revealed useful commonalities in the mutations that result in tumor formation and growth. Their findings suggest that such computational approaches could outperform existing profiling methods for other cancers and lead to more personalized treatments.
The study, published in the Proceedings of the National Academy of Sciences, was led by Ravi Radhakrishnan, Professor and chair of Penn Engineering’s Department of Bioengineering and professor in its Department of Chemical and Biomolecular Engineering, and Mark A. Lemmon, Professor of Pharmacology at Yale and co-director of Yale’s Cancer Biology Institute. The study’s first authors were Keshav Patil, a graduate student in Penn Engineering’s Department of Chemical and Biomolecular Engineering, along with Earl Joseph Jordan and Jin H. Park, then members of the Graduate Group in Biochemistry and Molecular Biology in Penn’s Perelman School of Medicine. Krishna Suresh, an undergraduate student in Radhakrishnan’s lab, Courtney M. Smith, a graduate student in Lemmon’s lab, and Abigail A. Lemmon, an undergraduate in Lemmon’s lab, contributed to the study. They collaborated with Yaël P. Mossé, Associate Professor of Pediatrics at Penn Medicine and in the division of oncology at CHOP.
“Some cancers rely on the aberrant activation of a single gene product for tumor initiation and progression,” says Radhakrishnan. “This unique mutational signature may hold the key to understanding which patients suffer from aggressive forms of the disease or for whom a given therapeutic drug may yield short- or long-term benefits. Yet, outside of a few commonly occurring ‘hotspot’ mutations, experimental studies of clinically observed mutations are not commonly pursued.”
In their study, the researchers compared some of these clinically observed mutations that occur in anaplastic lymphoma kinase (ALK), a known oncogenic driver in pediatric neuroblastoma, with “test” mutations computationally synthesized in their model. Combining their real and synthetic sets into a pool with an equal number of harmful and benign mutations, the researchers then used their model to predict which were which.
“We found that the computational methods discussed in our studies outperformed existing methods in the literature in predicting the activating status of the curated mutational set,” Jordan says.
“Moreover, the physics-based molecular dynamics method, leveraging the computational power of supercomputers to solve Newton’s equations for molecular systems, outperformed data-driven methods that rely on supervised machine learning,” Patil says.
One of the computational study’s findings was that two-thirds of cancer-driving ALK mutations involve the same general mechanism: the perturbation of hydrogen bonds in particular subdomains of the kinase, which activate their oncogenic properties.
“We should note that one limitation of our study is that we do not consider the clonal diversity of mutations in other genes in our predictions,” Park says. “This aspect is a subject of current research in the field of cancer, as evolutionary biology can play an important role in oncogene-driven tumors.”
“However,” Lemmon says, “by adopting a reductionist approach closely combining theoretical and experimental studies, we show that even by studying well-defined molecular and cellular systems, much can be gained in terms of a mechanism-based understanding of cellular decision-making processes and how they can go awry in diseases such as cancers.”
“Furthermore,” Mossé adds, “we hope to integrate this methodology into an in silico platform that can allow clinicians and researchers to design treatment in the most scientifically grounded, quantitative and patient-individualized fashion.”
This research was supported by funding from European Commission grant FP7-ICT-2011-9-600841 and National Institutes of Health grants R01 CA244660, R01 CA140198, U01 CA227550 and R35 GM122485. This work used the Extreme Science and Engineering Discovery Environment, which is supported by National Science Foundation grant ACI-1548562.
